
ALEXIS COULLOMB
ASSOCIATE INVESTIGATORS: JULIE BORDENAVE & NINA VERSTRAETE
Multi-omics study of spatial tumour microenvironment organization
Solid tumors are made of various cell types, such as tumor cells and pro- or anti-tumor immune cells, whose proportion and spatial organization play a decisive role in tumor progression or response to treatments. Moreover, numerous methods have been developed to produce maps of protein or RNA abundances at single cell resolution, such as multiplex immunofluorescence or cyclic RNA-FISH.
Our team develops tools to analyze the highly complex data generated by these methods, and we think that networks provide a powerful framework to analyze spatial omics experiments. The tysserand library can reconstruct spatial networks from spatially resolved omics experiments, and the mosna library (Multi Omics Spatial Networks Analysis) provides methods to analyze these spatial networks to study cell-cell interactions and find cellular neighborhoods of potential clinical interest.
Additionally, we are implementing FISH protocols to produce spatial omics data relative to immuno-oncology.
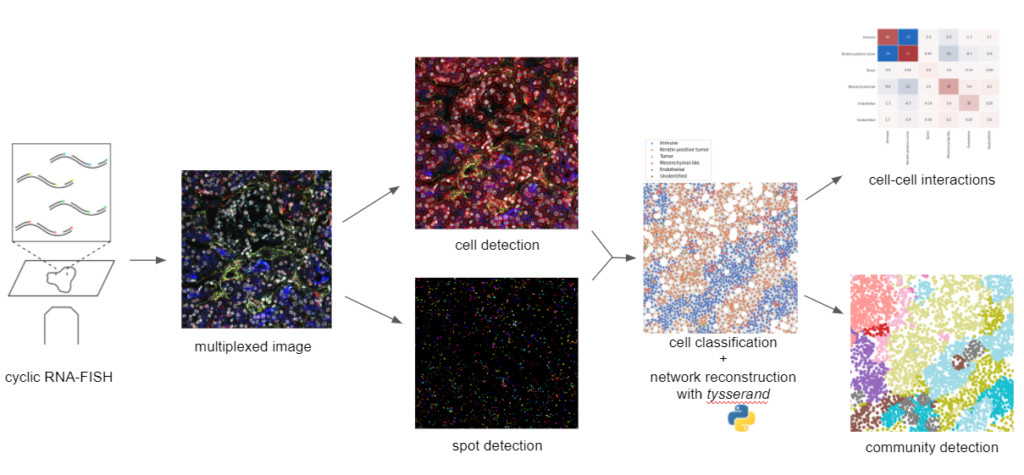
Single-cell spatial transcriptomics pipeline: cyclic RNA FISH consists of successive cycles of hybridization of fluorophore-conjugated probes to RNAs of interest, followed by hybridization for subsequent cycles. The resulting data is a multiplex image, with as many fluorescence channels as RNA species targeted by the FISH probes. After detection of individual cells by cell segmentation and RNA molecules by dot detection, this information is combined to define the phenotypes of individual cells, and to reconstruct the spatial tissue network with the tysserand library. This spatial network is then used to inspect cell-cell interactions and find spatial neighborhoods (or “niches”) of potential clinical interest.
Funding: Janssen
CRCT Chair of Informatics in Oncology (Fondation Toulouse Cancer Sante, Inserm et Pierre Fabre)
Pancaldi and Coullomb 2021
https://academic.oup.com/bioinformatics/advance-article-abstract/doi/10.1093/bioinformatics/btab490/6313163?redirectedFrom=fulltext
Github : https://github.com/VeraPancaldiLab/tysserand.

