
Molecular mechanisms and modulation of small GTPase activity in cancer
Stéphanie Cabantous
Most cellular functions depend on the fine coordination of signalling pathways defined by complex interaction networks. These protein-protein interactions (PPIs) are often context-specific, depending on the cellular environment, tissue type, and other dynamically changing phenotypical conditions. PPIs have crucial roles in network connectivity maintaining cancer cells growth, transmitting oncogenic signals and gaining hallmark features of cancer. Deciphering the variations in the protein interactome landscape is of crucial importance to a better understanding of tumorigenesis and the deregulation of cell signaling in cancer.
Methodology
We have developed tools based on split-GFP tripartite complementation that allow the analysis of protein-protein interactions at the single cell level (Cabantous, Nguyen et al. 2013). We have validated this approach for the analysis of the activation of small GTPases of the RAS superfamily in different human cell models (Cabantous, Nguyen et al. 2013). In this assay, two fragments of GFP (green fluorescent protein), strands s10 and s11 (commonly called GFP10 and GFP11), are fused with the GTPase and its corresponding effector domain, respectively. These fusion proteins (GFP10-GTPase and effector domain-GFP11) are co-expressed with the GFP1-9 detector fragment of GFP. Only when the GTPase is bound to GTP in the active conformation does association with the effector domain occur, bringing the GFP10 and GFP11 strands together and allowing complementation with GFP1-9 to reconstitute a fluorescent GFP (Figure 1). We have validated the specificity of the signal for the study of RHO and RAS GTPase activation (Koraichi, Gence et al. 2018) and have developed optimized cell models to perform measurements in a miniaturised format (Cabantous, WO2016207313).
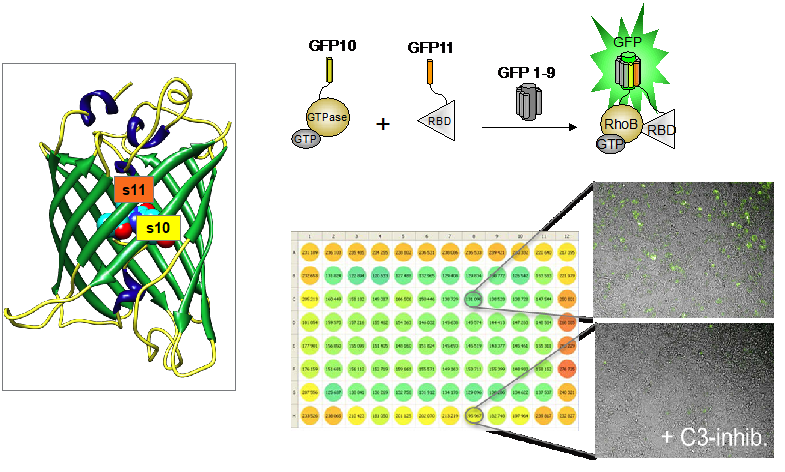
Figure 1. Modèles Split-GFP pour l’analyse des interactions entre protéines de petites GTPases et pour le criblage à haut débit d’inhibiteurs / Figure 1. Split-GFP models for the analysis of small GTPase protein interactions and for high-throughput screening of inhibitors
Axis 1 : Elucidating the molecular mechanisms of RhoB GTPase activity in targeted cancer therapies
Our main research activity concerns the identification of signalling pathways regulated by RHO small GTPases in lung oncogenesis in order to identify new actors of resistance in these cancers. We identify direct effectors of RHOB by high-throughput screening of human ORFeome libraries (collaboration Y. Jacob & Caroline Demeret, Institut Pasteur). These targets are then characterized for their impact on cell proliferation and on the modulation of cell survival pathways. The molecular and cellular characterization of the complex candidates is completed by mass spectrometry studies (Marie-Pierre Bousquet, IPBS, Toulouse), and by biophysical and structural analysis (Pédelacq Jean-Denis, IPBS). The combination of these data will be used to determine the architecture of these complexes to identify the sites of interactions in order to understand the functional consequences of these interactions, and to define potential regulators of these protein complexes that could serve as alternative pharmacological targets.
Axis 2 : Screening for inhibitors of RHO and RAS GTPases activity
An important application of the split-GFP biosensor models is the search for inhibitors of protein-protein interactions. Concerning small GTPases, the development of such molecules is still emerging or at the stage of in vitro or preclinical use, but could have applications in various diseases. In particular, our interest lies in the study of RHOB activation for which we have developed a screening model to identify specific inhibitors that could be used in endothelial cell models where RHOB is upregulated in response to inflammatory cytokines (Marcos-Ramiro, Garcia-Weber et al. 2016). Besides, we develop cell models to search for KRAS inhibitors in the context of lung adenocarcinoma (collaboration D. Santamaria).
Keywords:
- split-GFP,
- fluorescent proteins,
- small GTPases,
- RHO, RAS,
- cell signaling,
- high-throughput screening,
- protein-protein interaction.
Group members:
- Delphine Pagan, Assistant Engineer
- Carl Khawly, Master2 student.
Other team members involved:
- Gilles Favre, (PU, PH)
- Rémi Gence, Laboratory Engineer
- Béatrice Cena, Laboratory Engineer
Main external collaborations:
- Jean Denis Pédelacq, Structural Biology group, IPBS, Toulouse
- Marie Pierre Bousquet, Mass Spectrometry, IPBS, Toulouse
- Caroline Demeret, Interactomics of Viruses, Institut Pasteur
- Stanislas Faguer & Muriel Laffargue, Nephrology unit-CHU Rangueil, I2MC, Toulouse
- Manos Mavrakis, Institut Fresnel, Marseille
- David Santamaria, Institut de Salamanca, Espagne.
Selected publications :
Eur J Cell Biol. 2023 Dec;102(4):151355; PMID: 37639782
Castillo S, Gence R, Pagan D, Koraïchi F, Bouchenot C, Pons BJ, Boëlle B, Olichon A, Lajoie-Mazenc I, Favre G, Pédelacq JD, Cabantous S.
Visualizing the subcellular localization of RHOB-GTP and GTPase-Effector complexes using a split-GFP/nanobody labelling assay
Int J Mol Sci. 2019 Jul 15; 20 (14), 3479. DOI: 10.3390/ijms20143479.
Pedelacq JD, Cabantous S.
Development and Applications of Superfolder and Split Fluorescent Protein Detection Systems in Biology. Special Issue Imaging with Fluorescent Proteins.
Methods Mol Biol. 2019 2025:423-437. DOI: 10.1007/978-1-4939-9624-7_20
Pedelacq JD, Waldo GS, Cabantous S.
High-Throughput Protein-Protein Interaction Assays Using Tripartite Split-GFP Complementation.
J Cell Sci. 2018 Jan 1;131(1). DOI: 10.1242/jcs.210419
Koraïchi, F., Gence, R., Bouchenot, C., Grosjean, S., Lajoie-Mazenc, I., Favre, G., Cabantous,
S. High-content tripartite split-GFP cell-based assays to screen for modulators of small GTPase activation.
Sci Rep. 2013 Oct 4;3:2854. https://doi.org/10.1038/srep02854
Cabantous S*, Nguyen HB, Pedelacq JD, Koraïchi F, Chaudhary A, Ganguly K, Lockard MA, Favre G, Terwilliger TC, & Waldo GS* (*co-corresponding authors).
A new protein-protein interaction sensor based on tripartite split-GFP association.
Biotechniques 2010 Oct; 49(4):727-8, 730, 732 passim. doi: 10.2144/000113512. PubMed PMID: 20964633. DOI: 2144/000113512
Kaddoum L, Magdeleine E, Waldo GS, Joly E, Cabantous S.
One-step split GFP staining for sensitive protein detection and localization in mammalian cells.
